Translate this page into:
Gut Microbiota and the Ways to Correct it in Chronic Kidney Disease
Address for correspondence: Dr. Nikolay V. Sturov, Peoples’ Friendship University of Russia (RUDN University), Department of General Practice, 6 Miklukho-Maklaya Street, Moscow, 117198, Russian Federation. E-mail: sturov-nv@rudn.ru
-
Received: ,
Accepted: ,
This article was originally published by Wolters Kluwer - Medknow and was migrated to Scientific Scholar after the change of Publisher.
Abstract
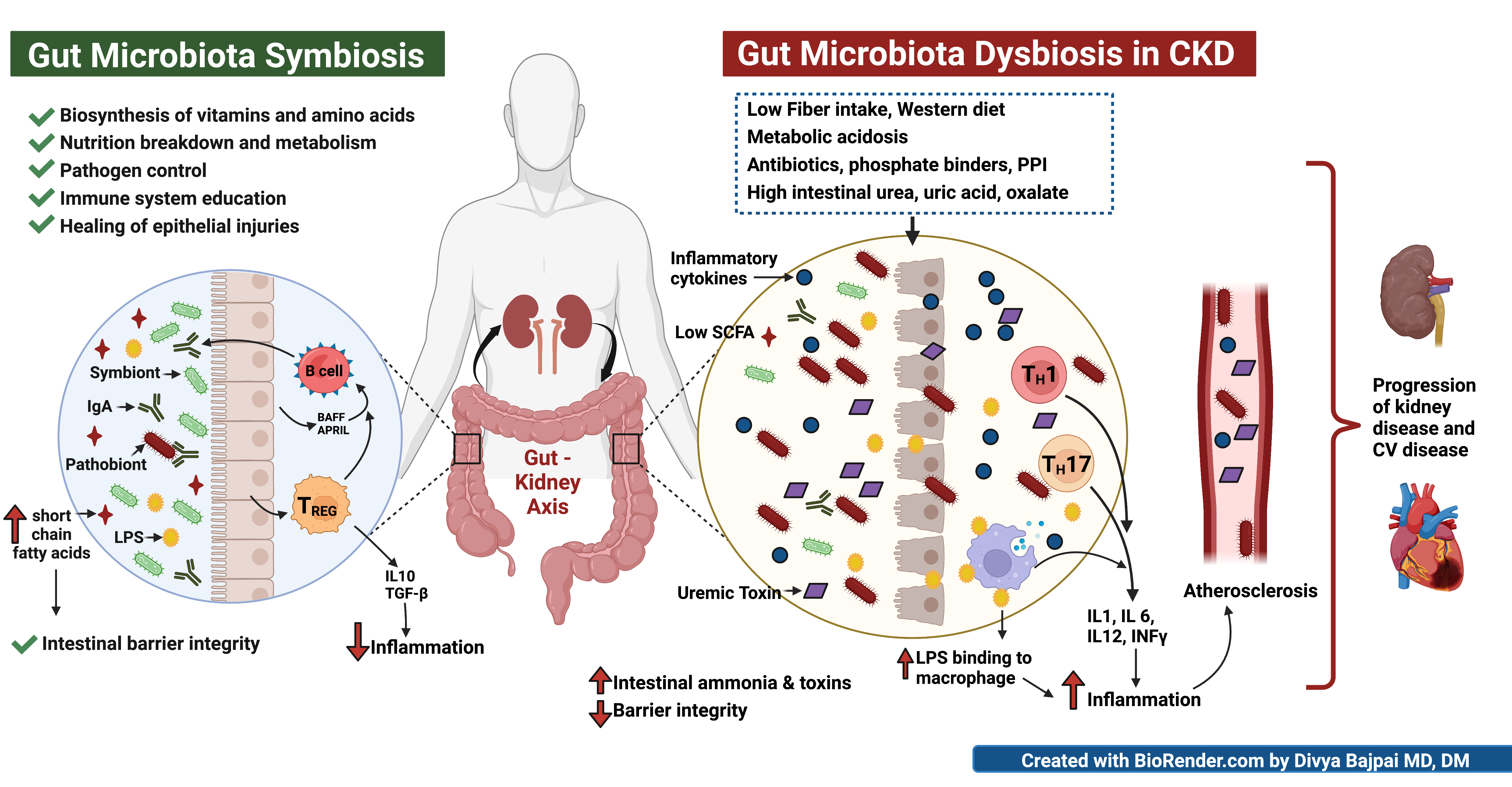
Approximately 13% of the Russian population suffers from chronic kidney disease (CKD). Such a high prevalence of the disease, as well as the complexity and high cost of renal replacement therapy, explain the need for developing and implementing new approaches to treat patients at the pre-dialysis stages. The data collected in recent decades highlight the importance of gut microbiota in the progression of CKD. This review provides information about the microbiota composition in healthy individuals and patients with CKD and discusses the mechanisms of interaction in the intestine–kidney system. The article also presents the specifics of the violation of gut microbiota (GM) and correction thereof in CKD.
Keywords
Chronic kidney disease
dysbiosis
enterotype
gut microbiota
intestine–kidney
Introduction
Chronic kidney disease (CKD) is an important medical and public health problem. According to several authors, the true epidemiology of this disease in the Russian Federation remains unknown due to the limited amount of data.[1-3] It is known that in the period from 2003 to 2013, the number of patients with CKD increased 2.2 times, and the average annual increase was approximately 9%.[1] Extrapolation of data from other countries suggests that the prevalence of CKD in the Russian Federation corresponds to that in the world (13.4%)[4,5] and is comparable with the figures obtained in many countries [Table 1]. Economic and social features (including the cost of treatment and the level of regional development) make the problem of CKD extremely relevant for domestic medicine.[3] Over the past 15 years, the average increase in the dialysis group among CKD patients was 10%, and 13% in 2018, but despite significant improvements in the availability of renal replacement therapy (RRT) in the Russian Federation, the availability of hemodialysis (HD) for the population is 2.5–7 times lower than that in the European Union.[3,6,7] This situation is aggravated by the fact that more than a quarter of patients seek specialized nephrological care at the stages when the possibility of administering nephroprotective therapy is missed and HD is required.[7] Although the impact of the new coronavirus infection on CKD has not yet been established, experts predict an increase in its incidence as up to 20% of hospitalized patients need RRT.[8,9]
| Country | Studies, year | CKD prevalence | |
|---|---|---|---|
| Stages 1-5 | Stages 3-5 | ||
| USA | NHANES, 1999-2006 | 15% | 8% |
| Netherlands | PREVEND, 2005 | 18% | No data |
| Spain | EPIRCE, 2005 | 13% | No data |
| China | Beijing study, 2008 | 14% | 7% |
| Japan | Imai et al., 2007 | No data | 19% |
| Australia | AusDiab, 2008 | 13% | 8% |
| Congo | Kinshasa study, 2009 | 12% | 8% |
CKD - Chronic kidney disease
In addition to the social burden associated with the disease, CKD has a significant impact on the healthcare system from an economic point of view. For example, the cost of kidney transplantation and RRT is 2%–3% of the national health expenditure in the Russian Federation. CKD is a comorbid disease in 16% of the working-age population of the country.[6] The data accumulated in recent decades emphasize the relationship of gut microbiota (GM) with the course of CKD. When entering the bloodstream, the toxins synthesized by GM in CKD patients contribute to a decrease in the glomerular filtration rate (GFR) and the progression of renal failure.[10] Moreover, there are no recommendations for the treatment of GM disorders in patients with CKD.
Composition of the Gut Microbiota
According to the Russian Industry Standard for the treatment of intestinal dysbiosis (OST 91500.11.0004-2003), the normal flora means the ratio of diverse populations of microorganisms of individual organs and systems that maintain the biochemical, metabolic, and immunological balance necessary to preserve human health.[11] GM is a complex community of microorganisms inhabiting the gastrointestinal tract (GIT) of a human: bacteria, viruses, prokaryotes, eukaryotes, and archaea.[12]
During prenatal development, the intestine is sterile, and its primary colonization occurs when passing through the birth canal.[13] It is known that the first colonizers of the GIT are aerophilous representatives of the Proteobacteria and Actinobacteria phyla, which form a favorable microenvironment for further colonization by representatives of Firmicutes and Bacteroides.[14] The microbial diversity of the intestine increases during the first 12 months of human life, and at about 2.5 years of age, the GM composition approaches that of adults.[15] In the course of life, the difference in the GM composition of a particular person from the average decreases, and the diversity between people, on the contrary, increases.[14]
In general, the total GM mass amounts to 1.5–2.0 kg, and the taxonomic diversity, once estimated at 500 species, is now more than 1500 species.[16,17] Because the human GIT is an extended structure that differs in morphological and physiological terms in different segments, the composition and functions of the microbiota also differ. For example, the jejunum is dominated by aerobic bacteria: enterobacteria, streptococci and staphylococci, lactobacilli, and yeast, while the anaerobic flora makes up a small part of the local diversity. In total, the jejunum contains up to 105 colony-forming units (hereinafter referred to as CFU) per 1 g of contents.
As we move from the ileum to the large intestine, not only does the total number of microorganisms increase (up to 109 CFU in the ileum and up to 1012 CFU in the large intestine) but the composition of the microbial community also changes, revealing the dominance of anaerobes in the large intestine.[18] In the large intestine, the bacteriodes of the families Prevotellaceae and Rikenellaceae predominate, and among the clostridiae, the family Lachnospiraceae dominates.[15] The composition of the gastrointestinal microbiota by segments is presented in Table 2.
| Microorganisms | Quantity, CFU/g of contents | |||
|---|---|---|---|---|
| Stomach | Jejunum | Ileum | Large intestine | |
| Total quantity | Up to 103 | Up to 105 | Up to 109 | Up to 1012 |
| Aerobes | ||||
| Family Enterobacteriaceae (gram-negative, nonsporeforming), of which (in total): | Up to 102 | Up to 103 | 102-107 | 104-1010 |
| E. coli, typical, lactose-negative, and hemolytic | none | none | none | 104-108 |
| Other potentially pathogenic enterobacteria | none | none | none | Up to 104 |
| Streptococci, including lactic acid (gram-positive, nonsporeforming) | Up to 103 | Up to 104 | 102-106 | 106-1010 |
| Staphylococci (gram-positive, nonsporeforming) | Up to 102 | Up to 103 | 102-106 | 104-109 |
| Lactobacilli (gram-positive, nonsporeforming) | Up to 103 | Up to 104 | 102-103 | 106-1010 |
| Yeast (gram-positive, sporeforming | Up to 103 | Up to 102 | 102-104 | 104-106 |
| Anaerobes | ||||
| Bacterioids (gram-negative, nonsporeforming) | rarely | 0-103 | 103-107 | 109-1012 |
| Bifidobacteria (gram-positive, nonsporeforming) | rarely | 0-104 | 103-109 | 104-1011 |
| Pepto-streptococci (gram-positive, non-sporeforming) | rarely | 0-103 | 102-106 | 1010-1012 |
| Clostridia (gram-positive, spore-forming) | rarely | rarely | 102-104 | 106-1011 (according to other sources, up to 105) |
| Eubacteria (gram-negative and gram-positive, spore-forming and non-sporeforming) | rarely | rarely | rarely | 109-1012 |
For a long time, the composition of the microbial contents of the intestine was analyzed only with the help of fecal microbiological studies. Currently, metagenomic sequencing of the 16S rRNA gene from amplified bacterial nucleic acid isolated from feces is used.[15] Thanks to the results of the Human Microbiome Project (HMP) and Metagenomics of Human Intestinal Tract (MetaHIT) project aimed at studying the composition of normal GM in American and European populations, current ideas about humans, by analogy with blood groups, were supplemented by the classification of people according to the GM composition [Table 3].
| Nomenclature of bacteria | HPM (American) | MetaHIT (European) |
|---|---|---|
| Bacteroidetes phylum: | ||
| Bacteroides spp | 51.1% | 21.8% |
| Alisitpes spp | 12.3% | 8.6% |
| Prevotella spp | 5.7% | 11.8% |
| Other species | 6.6% | 3.6% |
| Firmicutes phylum: | ||
| Eubacterium spp | 6.9% | 14.7% |
| Ruminococcus spp | 3.7% | 6.0% |
| Faecalibacterium spp | 3.5% | 5.7% |
| Dialister spp | 2.3% | 4.0% |
| Roseburia spp | 1.5% | 3.9% |
| Butyrivibrio spp | 1.5% | 3.0% |
| Other species | 2.6% | 6.0% |
| Protobacteria phylum: | ||
| Sutterella spp | 1.1% | 0.6% |
| Escherichia spp | 0.8% | 1.0% |
| Other species | 0.2% | 1.0% |
| Verrucomicrobia phylum: | ||
| Akkermansia spp | 0.9% | 2.3% |
| Actinobacteria phylum: | ||
| Bifidobacterium spp | 0.4% | 2.1% |
| Other phyla: | ||
| Other species | 0.3% | 0.5% |
GM growth is limited by the intestinal barrier, which includes physical (epithelial and mucosal layers), biochemical (enzymes and antimicrobial proteins), and immunological (epithelial immune cells and IgA) components.[15]
The variety of GM representatives increases relatively quickly by the age of 25–30 and then gradually reaches a peak at the age of 50, followed by a gradual decline.[16] After age 65, the microbial diversity changes—the number of Bacteroidetes phyla and Clostridium increases.[15] In addition to age, the diversity of the composition is influenced by gender (in women, it is significantly higher than in men of the same age), hematocrit, blood plasma lipid profile, several secreted proteins, and peptides.[16] Among the dietary factors that affect the GM composition, some researchers distinguish the qualitative and quantitative composition of the diet, the content of sugar, polyphenols, fat content of the consumed milk, and other factors.[16,20] Glycosylation of mucus and mucin plays a key role in the formation of the microbiota. In the case of dietary fiber deficiency, mucosal erosion is associated with the switching of GM to the use of secreted mucins as a source of nutrients.[15] The issue of iatrogenic dysbiosis also deserves special attention; for example, changes in GM have been proven with the use of proton pump blockers, metformin, statins, laxatives, neuroleptics, antidepressants, and antibiotics.[16,20]
Researchers pay great attention to studying the relationship of changes in GM and the associated metabolites with the stages of CKD. Thus, in the early stages of CKD, a decrease in the number of Bacteroides eggerthii is detected. By reducing the number of Prevotella sp. 885, which correlates with the excretion of urea in the daily urine, signs of progression of this disease can be detected. In the late stages of CKD, the increase in serum lipopolysaccharides (LPS) is partly due to an increase in the number of Escherichia coli and other representatives of the Enterobacteriaceae family in the large intestine. A significant decrease in the level of propionic acid also indicates a late stage of CKD, and its absence is a reliable sign of a serious condition of a patient.[21]
Enterotypes of the Gut Microbiota
According to the results of 2007–2019 studies, there are three enterotypes of gut microbial communities.[22] Enterotype 1 is characterized by the predominance of Bacteroides spp., enterotype 2 by Prevotella spp., and enterotype 3 by representatives of the Firmicutes phylum, including species such as Ruminococcus and Faecalibacterium.[19] The researchers found that the enterotypes differ in the ability to process incoming food components, the synthesis of vitamins, etc., It was noted that enterotype 1 is more common in people who prefer to consume a large amount of proteins and fats of animal origin, and enterotype 2 in people whose diet is predominantly plant-based. The formation of enterotype 3 is promoted by a diet rich in carbohydrates.[23-25]
Another aspect that influences the GM composition is the ethnic origin of a person. In 2020, the presence of population and continental specificity of enterotypes was proved due to the fact that the area of residence determines the dietary pattern. It has been established that such differences are caused by the genetic characteristics of certain ethnic groups under conditions of homogeneous environmental factors.[23-25]
Gut Microbiota in CKD
The data accumulated to date on changes in the microbiota in CKD are characterized by high heterogeneity. The predominant number of studies was conducted on patients with stage 5 CKD. The number of studies on pre-dialysis patients is significantly less.
In several studies, the results of quantitative changes coincide. Thus, the authors clearly show a decrease in the number of bacteria from the families Bifidobacteriaceae, Lactobacillaceae, and Prevotellaceae. Members of the families Micrococcaceae, Clostridiaceae, Peptostreptococcaceae, and Pseudomonadaceae are reduced in patients at different stages of CKD in different countries. Regarding the families Lachnospiraceae and Enterobacteriaceae, the authors demonstrate conflicting results. With regard to representatives of other families, comparison of the results is not possible as the analysis of each of these families of bacteria is presented in only one work. It is important to note that two published studies in patients with pre-dialysis stages of CKD did not reveal quantitative differences in the composition of the intestinal microbiota compared with healthy controls [Table 4].
| Nomenclature of bacteria | Stages of CKD | Increase | Decrease | Country | References |
|---|---|---|---|---|---|
| Actinobacteria | |||||
| Atopobiaceae | 4-5 | Yes | China | [26] | |
| Beutenbergiaceae | 5 | Yes | USA | [27] | |
| Bifidobacteriaceae | 5 | Yes | Taiwan | [28] | |
| 1-5 | Yes | Taiwan | [21] | ||
| Brachybacterium | 5 | Yes | USA | [29] | |
| Cellulomonadaceae | 5 | Yes | USA | [27] | |
| Coriobacteriaceae | 4-5 | Yes | China | [30] | |
| Corynebacteriaceae | 4-5 | Yes | China | [26] | |
| Dermabacteraceae | 5 | Yes | USA | [27] | |
| Micrococcaceae | 5 | Yes | USA | [27] | |
| 4-5 | Yes | China | [26] | ||
| Nesterenkonia | 5 | Yes | USA | [29] | |
| Bacteroidetes | |||||
| Prevotellaceae | 5 | Yes | USA | [27] | |
| 5 | Yes | Taiwan | [21] | ||
| 5 | Yes | China | [31] | ||
| Tannerellaceae | 5 | Yes | China | [30] | |
| Firmicutes | |||||
| Acidaminococcaceae | 5 | Yes | China | [26] | |
| Catabacter | 5 | Yes | USA | [27,29] | |
| Catenibacterium | 5 | Yes | USA | [29] | |
| Clostridiaceae | 5 | Yes | USA | [27] | |
| 5 | Yes | China | [26] | ||
| Yes | Austria | [32] | |||
| Coprobacillaceae | 5 | Yes | USA | [27] | |
| Enterococcaceae | 5 | Yes | Taiwan | [28] | |
| Erysipelotrichaceae | 5 | Yes | Taiwan | [21] | |
| Lachnospiraceae | 5 | Yes | China | [26] | |
| 5 | Yes | Austria | [32] | ||
| 5 | Yes | China | [31] | ||
| Lactobacillaceae | 5 | Yes | USA | [27] | |
| 5 | Yes | Taiwan | [28] | ||
| 1-5 | Yes | Taiwan | [21] | ||
| Peptococcaceae | 4-5 | Yes | China | [26] | |
| Peptostreptococcaceae | 5 | Yes | USA | [29] | |
| 5 | Yes | China | [30] | ||
| Proteobacteria | |||||
| Alcaligenaceae | 5 | Yes | USA | [27] | |
| Alteromonas | 5 | Yes | USA | [27,29] | |
| Enterobacteriaceae | 5 | Yes | USA | [29] | |
| 5 | Yes | Taiwan | [28] | ||
| Halomonadaceae | 5 | Yes | USA | [27,29] | |
| Methylococcaceae | 5 | Yes | USA | [27,29] | |
| Moraxellaceae | 5 | Yes | USA | [27,29] | |
| Polyangiaceae | 5 | Yes | USA | [27,29] | |
| Pseudomonadaceae | 5 | Yes | USA | [27,29] | |
| 5 | Yes | Taiwan | [28] | ||
| Thiothrix | 5 | Yes | USA | [27,29] | |
| Xanthomonadaceae | 5 | Yes | USA | [27] | |
| Verrucomicrobia | |||||
| Verrucomicrobiaceae | 5 | Yes | USA | [27] | |
| All families | 3-4 | No difference | Brazil | [33] | |
| 1--5 | Belgium | [34] | |||
Presumably, the reason for the high variability of the obtained data is several factors. As shown above, in healthy individuals, the composition of the intestinal microbiota is determined by sex, age, type of diet taken by pharmacotherapy, and biochemical processes in enterocytes. It seems appropriate to conduct a systematic review of the composition of the intestinal microbiota in control groups and evaluate quantitative changes in the light of demographic and environmental data on patients, but such an analysis was not carried out in the reviewed publications. Conducting studies with large numbers of patients, dividing into groups not only by the stage of CKD but also by other factors, will expand the understanding of the mutual influence of external and internal environmental factors, CKD, and the quantitative diversity of the intestinal microbiota in patients. Greater involvement of patients at the pre-dialysis stages and long-term observational studies are also important for understanding the dynamics of GM composition.
The concept of the microbiota as a separate organ, which emerged in the last decade, was naturally advanced to the intestine–kidney system.[35] GM can affect both the function of the intestinal wall and various human organs (kidneys, adipose tissue, hypothalamus–pituitary–kidney system, etc.) through low-molecular-weight mediators and metabolites that enter the blood. In 2016, researchers noted that uremic toxins of intestinal origin play a significant role in the progression of CKD.[10,36,37]
Under the influence of increased concentrations of urea in the microcirculatory bed, the permeability of enterocytes increases due to its direct impact on membrane proteins. As a result, urea enters the lumen of the intestine. In response, intestinal bacteria begin to produce urease, under the influence of which urea breaks down to ammonium hydroxide. Ammonium hydroxide is absorbed through the intestinal wall into the blood along with other toxic waste products of GM, which leads to chronic intoxication and encephalopathy.[38-41]
In the late stages of CKD, a deficiency of Lactobacillus spp. and a decrease in the number of Bifidobacterium spp. and E. coli is observed in the large intestine with a parallel increase in the number of enteropathogenic strains. The number of enterobacteria of Enterobacter spp. and Citrobacter spp. also increases. The relationship between the level of urea in the blood and the amount of bacteroids in the feces that produce urease has been established. In turn, E. coli and Lactobacillus spp. are involved in the utilization of ammonia, playing their role in maintaining the integrity of the intestinal barrier. A decrease in E. coli and Lactobacillus spp. is followed by an increase in the amount of ammonia entering the bloodstream.[42-44]
The relation between intestinal dysbiosis and endotoxemia-mediated inflammation was demonstrated in a pilot study comparing the blood microbiome profile of patients with CKD and a control group with the help of 16S ribosomal RNA sequencing. As a result of metagenomic studies, the possibility of systemic bacteremia was confirmed. According to the results of the conducted studies, it was also confirmed that the GIT in CKD is a source of microorganisms in the bloodstream, leading to systemic inflammation and sepsis in HD patients.[37,45,46]
In the case of azotemia, the function of pancreatic beta cells is compromised and type 2 diabetes mellitus progresses in patients with CKD.[47,48] It is known that in azotemia, the process of protein carbamylation occurring in the patient’s body under the influence of excess urea contributes to the development of atherosclerosis and ultimately leads to an increase in the mortality of CKD patients. The authors of the study concluded that the plasma level of protein-bound homocitrulline (PBHCit), which results from carbamylation, is a predictor of increased cardiovascular risk in patients with stage 5 CKD, supporting the relationship between uremia, inflammation, and atherosclerosis.[49]
Uremic toxins include p-cresyl sulfate, indoxyl sulfate, cresol, and trimethylamine-N-oxide (TMAO), the levels of which increase in blood serum as kidney failure progresses. These toxins increase the permeability of the intestinal wall.[50-54] By their nature, these uremic toxins are products of protein metabolism; therefore, a low-protein diet should be considered the most important method of GM correction in CKD.[52,53,55]
In patients with CKD, GM-synthesized toxins when they enter the bloodstream contribute to a decrease in the GFR and the progression of renal failure, which was demonstrated through the example of the tryptophan amino acid.[56] GM-produced toxins that enter the bloodstream also affect the activation of serum protein kinases, increasing the calcification of the arteries mediated by these enzymes.[57-59]
GM Correction in CKD
In connection with the described role of GM in the pathogenesis of CKD, the need to study the ways to correct is obvious. GM correction in CKD should be carried out to reduce the production of uremic and intestinal toxins and prevent metabolic (insulin resistance, vascular calcification) and other disorders.[60] A low-protein diet, adsorbents, prebiotics, and probiotics can be used to correct GM. In case of successful correction, the intestinal wall permeability improves, the synthesis of intestinal toxins reduces, and the microflora in the large intestine normalizes.[61-63] Probiotics (bifidobacteria, lactobacteria) reduce the concentration of nephrotoxic products in the intestine, improve the intestinal epithelium, inhibit apoptosis of enterocytes, and normalize the barrier function of the intestinal wall.[64-66] Prebiotics, which are non-digestible components of food, also reduce the concentration of toxic products in the intestine and the severity of local inflammation caused by the toxic effect of urea on the intestinal wall, have a positive effect on human health, creating a microenvironment that stimulates the growth of useful components of the microbiota. Fructooligosaccharides, galactooligosaccharides, xylooligosaccharides, inulin, and pyrodextrins are widely used as such additives.[67-70]
Conclusive Statement
Thus, the study of the GM specifics and the nature of changes in its composition depending on the severity of CKD support the development of new approaches to the treatment of this category of patients. The correction of GM in CKD patients can be carried out to improve kidney function, the quality of life, and reduce the mortality rate.
Gratitude: The work was carried out within the framework of research work No. 033802-0-000 “Study of intestinal microflora and methods of its correction in patients with kidney and urinary tract diseases.”
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
References
- Chronic kidney disease: Prevalence and risk factors (literature review) Health Risk Anal. 2020;2:164-74.
- [Google Scholar]
- Chronic Kidney Disease: Selected Chapters of Nephrology. 2017. Moscow: GEOTAR-Media; :507. Available from: https://bookmos.ru/components/com_jshopping/files/img_products/hronicheskaya-bolezn-pochek-izbrannye- glavy-nefrologii-2017-978-5-9704-4192-3.pdf
- [Google Scholar]
- Preventive care for patients with chronic kidney disease in the Russian Federation: An analytical review of the prevalence and existing programs (review) Saratov Sci Med J. 2019;15:24-8.
- [Google Scholar]
- 2019. Ministry of Health of the Russian Federation. Chronic kidney disease (CKD). Clinical recommendations. (In Russ.) Available from: https://nonr.ru/?p=4092
- Replacement therapy of terminal chronic renal failure in the Russian federation in 2010-2015. Report on the data of the All-Russian register of renal replacement therapy of the Russian dialysis society. Nephrol Dial. 2017;19:2-94.
- [Google Scholar]
- Chronic kidney disease: The renoprotective therapy effect on patients life quality within the framework of health system standardization. Probl Stand Healthcare 2019:58-62. doi: 10.26347/1607-2502201905-06058-062
- [Google Scholar]
- Coronavirus COVID-19: Official information about coronavirus in Russia on the portal-stopkoronavirus.rf. Available from: https://стопкоронавирус.рф
- COVID-19-associated acute kidney injury: Consensus report of the 25th acute disease quality initiative (ADQI) workgroup. Nat Rev Nephrol. 2020;16:747-64.
- [Google Scholar]
- Minireview: Gut microbiota: The neglected endocrine organ. Mol Endocrinol. 2014;28:1221-38.
- [Google Scholar]
- OST 91500.11.0004-2003 Protocol of patient management. Intestinal dysbacteriosis, OST (Industry Standard) No. 91500.11.0004-2003 of June 09, 2003, approved by Order of the Ministry of Health of the Russian Federation No. 231 of June 9, 2003. Available from: https://docs.cntd.ru/document/1200119089
- Maternal modifiers of the infant gut microbiota: Metabolic consequences. J Endocrinol. 2017;235:R1-12.
- [Google Scholar]
- The first microbial colonizers of the human gut: Composition, activities, and health implications of the infant gut microbiota. Microbiol Mol Biol Rev. 2017;81:e00036-17.
- [Google Scholar]
- Microbiota of Various Loci of the Organism. 2017. Moscow: Russian Academy of Sciences; :38. Available from: elibrary.ru/item.asp?id=32753482
- [Google Scholar]
- Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science. 2016;352:565-9.
- [Google Scholar]
- Microbiota. 2019. Moscow: Media Sphere Publishing House; :255. Available from: https://www.elibrary.ru/item.asp?id=38244535
- [Google Scholar]
- Intestinal microflora part I – general overview. Folia Gastroenterol Hepatol. 2008;6:150-60.
- [Google Scholar]
- The gut microbiota and metabolic disease: Current understanding and future perspectives. J Intern Med. 2016;280:339-49.
- [Google Scholar]
- Gut microbiota composition is associated with polypharmacy in elderly hospitalized patients. Sci Rep. 2017;7:11102. doi: 10.1038/s41598-017-10734-y
- [Google Scholar]
- Integrative metagenomic and metabolomic analyses reveal severity-specific signatures of gut microbiota in chronic kidney disease. Theranostics. 2020;10:5398-411.
- [Google Scholar]
- A guide to enterotypes across the human body: Meta-analysis of microbial community structures in human microbiome datasets. PLoS Comput Biol. 2013;9:e1002863. doi: 10.1371/journal.pcbi. 1002863
- [Google Scholar]
- Microbiome and health implications for ethnic minorities after enforced lifestyle changes. Nat Med. 2020;26:1089-95.
- [Google Scholar]
- Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat Med. 2018;24:1526-31.
- [Google Scholar]
- Ethnic differences shape the alpha but not beta diversity of gut microbiota from school children in the absence of environmental differences. Microorganisms. 2020;8:254. doi: 10.3390/microorganisms8020254
- [Google Scholar]
- Impaired renal function and dysbiosis of gut microbiota contribute to increased trimethylamine-N-oxide in chronic kidney disease patients. Sci Rep. 2017;7(1):1-12. doi: 10.1038/s41598-017-01387-y
- [Google Scholar]
- Expansion of urease- and uricase-containing, indole- and p-cresol-forming and contraction of short-chain fatty acid-producing intestinal microbiota in ESRD. Am J Nephrol. 2014;39:230-7.
- [Google Scholar]
- Real-time PCR analysis of the intestinal microbiotas in peritoneal dialysis patients. Appl Environ Microbiol. 2012;78:1107-12.
- [Google Scholar]
- Chronic kidney disease alters intestinal microbial flora. Kidney Int. 2013;83:308-15.
- [Google Scholar]
- Altered gut microbiota and microbial biomarkers associated with chronic kidney disease. Microbiologyopen. 2019;8:e00678.
- [Google Scholar]
- Alteration of the gut microbiota in Chinese population with chronic kidney disease. Sci Rep. 2017;7:1-10.
- [Google Scholar]
- Structural and functional differences in gut microbiome composition in patients undergoing haemodialysis or peritoneal dialysis. Sci Rep. 2017;7:1-10.
- [Google Scholar]
- Is there interaction between gut microbial profile and cardiovascular risk in chronic kidney disease patients? Future Microbiol. 2015;10:517-26.
- [Google Scholar]
- Gut microbiome profiling uncovers a lower abundance of butyricicoccus in advanced stages of chronic kidney disease. J Pers Med. 2021;11:1118. doi: 10.3390/jpm11111118
- [Google Scholar]
- Gut microbiome in chronic kidney disease. Curr Hypertens Rep. 2017;19:29. doi: 10.1007/s11906-017-0727-0
- [Google Scholar]
- Chronic kidney disease: Pathophysiological role of dysbiosis of intestine and renoprotective effectiveness of interventions concerning its modulation. Med J. 2016;22:157-62.
- [Google Scholar]
- Altered intestinal microbial flora and impaired epithelial barrier structure and function in CKD: The nature, mechanisms, consequences and potential treatment. Nephrol Dial Transplant. 2016;31:737-46.
- [Google Scholar]
- Urea and impairment of the Gut-Kidney axis in chronic kidney disease. G Ital Nefrol. 2017;34:2017-vol6.
- [Google Scholar]
- The leaky gut and altered microbiome in chronic kidney disease. J Ren Nutr. 2017;27:458-61.
- [Google Scholar]
- Chronic kidney disease causes disruption of gastric and small intestinal epithelial tight junction. Am J Nephrol. 2013;38:99-103.
- [Google Scholar]
- The role of the intestinal microbiota in the course of chronic kidney disease. 2016. Gastroenterol St. Petersburg. :3-4. M2b Available from: https://www.elibrary.ru/item.asp?id=28297779
- [Google Scholar]
- Blood microbiome profile in CKD: A pilot study. Clin J Am Soc Nephrol. 2019;14:692-701.
- [Google Scholar]
- Uremic toxins in blood of end stage renal disease patients with dysbiosis of digestive tract. Experimental Clin Urol. 2014;2:94-7.
- [Google Scholar]
- Urea-induced ROS generation causes insulin resistance in mice with chronic renal failure. J Clin Invest. 2010;120:203-13.
- [Google Scholar]
- Urea impairs b cell glycolysis and insulin secretion in chronic kidney disease. J Clin Invest. 2016;126:3598-612.
- [Google Scholar]
- Protein carbamylation predicts mortality in ESRD. J Am Soc Nephrol. 2013;24:853-61.
- [Google Scholar]
- The uremic toxicity of indoxyl sulfate and p-Cresyl sulfate: A systematic review. J Am Soc Nephrol. 2014;25:1897-907.
- [Google Scholar]
- Gut microbiota-dependent trimethylamine N-oxide (TMAO) pathway contributes to both development of renal insufficiency and mortality risk in chronic kidney disease. Circ Res. 2014;116:448-55.
- [Google Scholar]
- Trimethylamine N-oxide from gut microbiota in chronic kidney disease patients: Focus on diet. J Ren Nutr. 2015;25:459-65.
- [Google Scholar]
- Disorder of gut amino acids metabolism during CKD progression is related with gut microbiota dysbiosis and metagenome change. J Pharm Biomed Anal. 2018;149:425-35.
- [Google Scholar]
- The role of gut microbiota and diet on uremic retention solutes production in the context of chronic kidney disease. Toxins (Basel). 2018;10:155. doi: 10.3390/toxins10040155
- [Google Scholar]
- The relationship between blood metabolites of the tryptophan pathway and kidney function: A bidirectional Mendelian randomization analysis. Sci Rep. 2020;10:12675. doi: 10.1038/s41598-020-69559-x
- [Google Scholar]
- Protein-bound uremic toxins.new targets to prevent insulin resistance and dysmetabolism in patients with chronic kidney disease. J Ren Nutr. 2013;23:464-6.
- [Google Scholar]
- Contributory role of gut microbiota and their metabolites toward cardiovascular complications in chronic kidney disease. Semin Nephrol. 2018;38:193-205.
- [Google Scholar]
- Indoxyl sulfate and p-Cresyl sulfate promote vascular calcification and associate with glucose intolerance. J Am Soc Nephrol. 2019;30:751-66.
- [Google Scholar]
- Intestinal microbiota and inflammation in chronic kidney disease. Lechaschi Vrach J. 2020;1:32-5.
- [Google Scholar]
- Role of neurochemicals in the interaction between the microbiota and the immune and the nervous system of the host organism. Probiotics Antimicrob Proteins. 2017;9:215-34.
- [Google Scholar]
- The use of enterosorbents in patients with chronic kidney disease at predialysis stage. Farmateka. 2016;6:76-83.
- [Google Scholar]
- Correction of intestinal microbiocenosis as a preventive measure for progression of chronic kidney disease. Meditsinskiy sovet Medical Council 2018:84-9. doi: 10.21518/2079-701X-2018-14-84-89
- [Google Scholar]
- Biomodulation of the toxic and nutritional effects of small bowel bacterial overgrowth in end-stage kidney disease using freeze-dried lactobacillus acidophilus. Miner Electrolyte Metab. 1996;22:92-6.
- [Google Scholar]
- Soluble proteins produced by probiotic bacteria regulate intestinal epithelial cell survival and growth. Gastroenterology. 2007;132:562-75.
- [Google Scholar]
- Probiotic lactobacilli and VSL#3 induce enterocyte b-defensin 2. Clin Exp Immunol. 2008;151:528-35.
- [Google Scholar]
- Gut microbiota and chronic kidney disease: Implications for novel mechanistic insights and therapeutic strategies. Int Urol Nephrol. 2018;50:289-99.
- [Google Scholar]
- Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia. 2007;50:2374-83.
- [Google Scholar]
- P-Cresyl sulfate serum concentrations in haemodialysis patients are reduced by the prebiotic oligofructose-enriched inulin. Nephrol Dial Transplant. 2010;25:219-24.
- [Google Scholar]
- Foods with added fiber lower serum creatinine levels in patients with chronic kidney disease. J Ren Nutr. 2013;23:e29-32.
- [Google Scholar]







